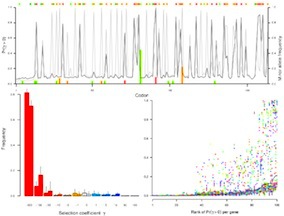
gammaMap
gammaMap is a program for detecting natural selection in sequence data from one or more species. It is based on a combined population genetics-phylogenetics model of selection. gammaMap estimates the distribution of selection coefficients, and allows localization of the signal of selection using a Bayesian sliding window approach. The signature of selection is detected from the contrast in the dN/dS ratio within and between species. The method and modelling approach are described in
Wilson, D. J., R. D. Hernandez, P. Andolfatto and M. Przeworski (2011)
A population genetics-phylogenetics approach to inferring natural selection in coding sequences.
PLoS Genetics 7: e1002395. (abstract
pdf)
A ready-to-use image is available for Docker and Singularity. Older platform-specific executables ares available to download from code.google.com/p/gcat-project/. Source code is available from the GitHub repository.